Synthetic biology and biomanufacturing automation in biofoundries
Biomanufacturing is harnessing synthetic biology to develop faster, more predictable, novel routes for fine and speciality chemicals production to deliver new chemical diversity towards scale-up and industrial manufacturing. Our group has contributed to developing design tools for metabolic engineering in the Design-Build-Test-Learn cycle. RetroPath2.0 (Met Eng, 2018) is an updated retrosynthesis framework with the ability of predicting de novo reactions by replacing native enzymatic reactions by generic reaction rules, expanding in that way natural chemical diversity. Such variable representation of reactions mimics the natural capabilities of enzymes that can promiscuously accept multiple substrates, allowing fine tuning of the pathway. The tool has been integrated with Selenzyme (Bioinformatics, 2018), an online enzyme selection tool for metabolic pathway design focusing on reaction rules.
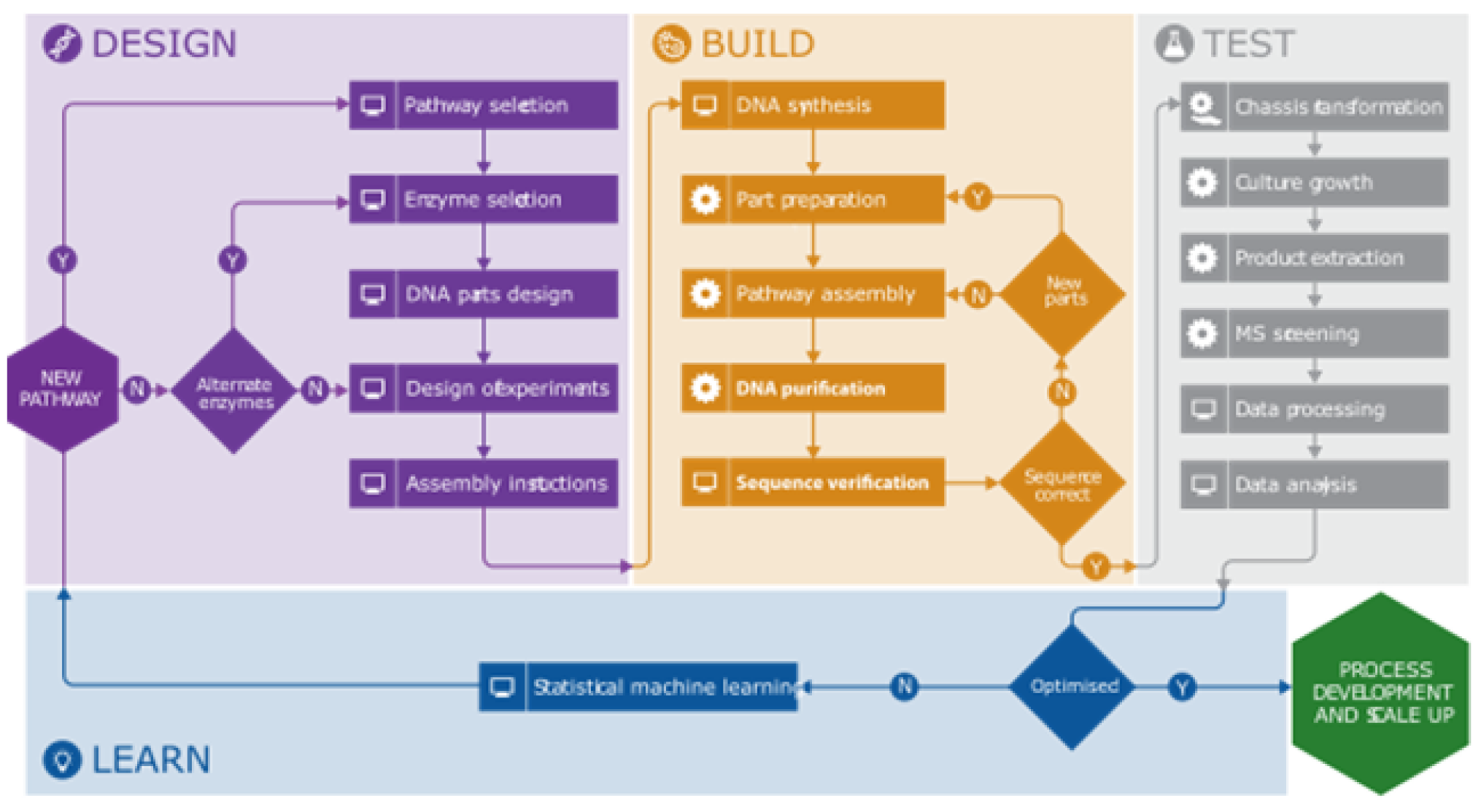
Those pathway Design tools were integrated into the Synbiochem pipeline (Figure 3). Reusable DNA parts are combined in silico into large combinatorial libraries of pathway designs, which are statistically reduced using a design of experiments (DoE) workflow to smaller representative libraries. The pipeline then proceeds to the Build (robotics platform) and Test (metabolomics platform) stages. The pipeline has been applied to produce the flavonoid pinocembrin in Escherichia coli, successfully establishing a production pathway improved by 500-fold, with competitive titers up to 484 mg L−1 (Synth Biol, 2020). The further application of the pipeline to optimize an alkaloids pathway demonstrated how it could facilitate the rapid optimization of microbial strains for production of any chemical compound of interest (Comm Biol, 2018).
Automation of the synthetic biology biomanufacturing pipeline. The automation of the different steps of the Design-Build-Test-Learn cycle through the integration of experimental data and traceability allows the rapid prototyping of production strains.

